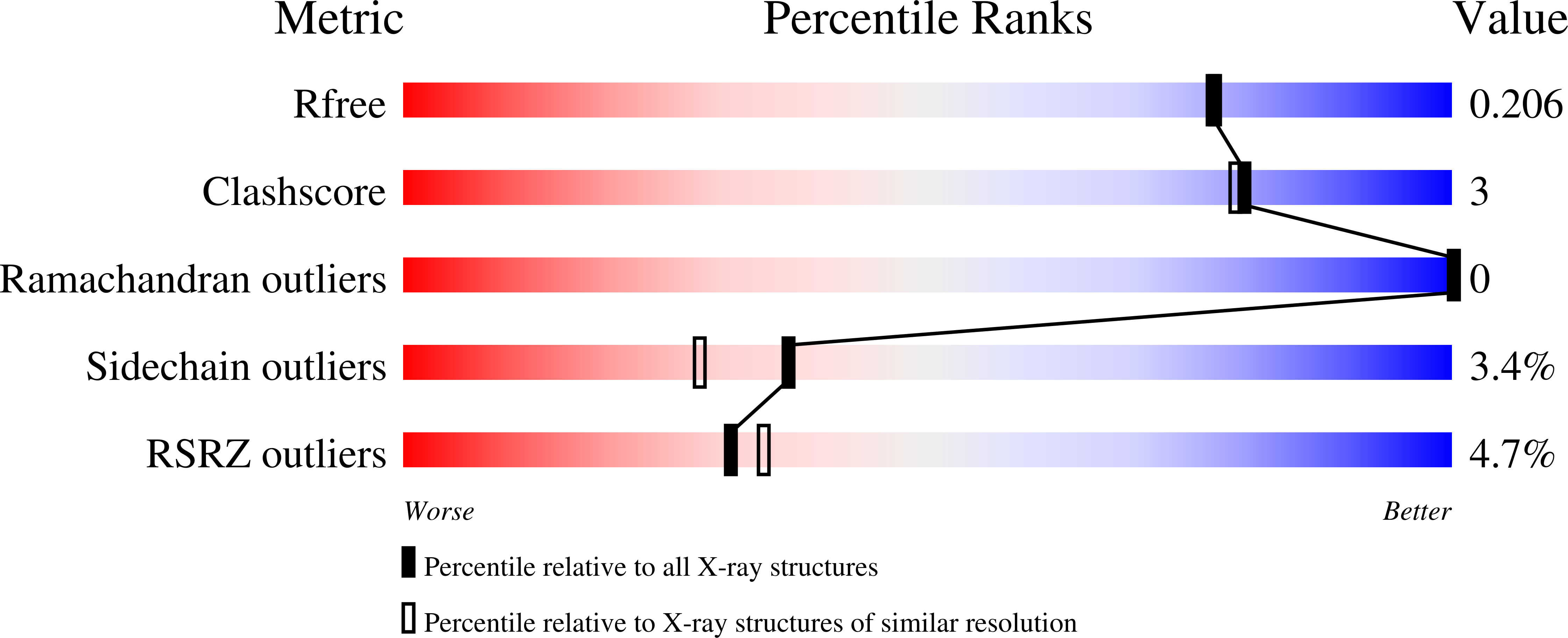
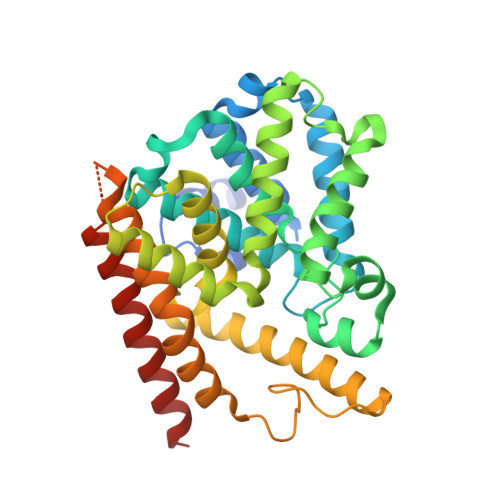
Small-molecule phosphodiesterase probes: discovery of potent and selective CNS-penetrable quinazoline inhibitors of PDE1
Humphrey, J.M., Yang, E.X., Am Ende, C.W., Arnold, E.P., Head, J.L., Jenkinsonb, S., Lebel, L.A., Liras, S., Pandit, J., Samas, B., Vajdos, F., Simons, S.P., Evdokimova, A., Mansour, M., Menniti, F.S.(2014) Medchemcomm